생물정보
데이터의 생산·분석·관리부터 교육까지 연구자가 데이터를 효과적으로활용할 수 있도록 지원하는 통합 생물정보 서비스를 제공합니다.
생물정보 기술을 기반으로 연구자들이 보다 정밀하고 효율적으로 데이터를 분석하고 활용할 수 있도록 지원합니다.NGS 기반 유전체, 전사체, 단백체 등 멀티오믹스 데이터 분석 서비스를 제공하며, 연구 목적에 맞춘 맞춤형 서비스를 통해 생물정보 연구의 정밀도를 높입니다.
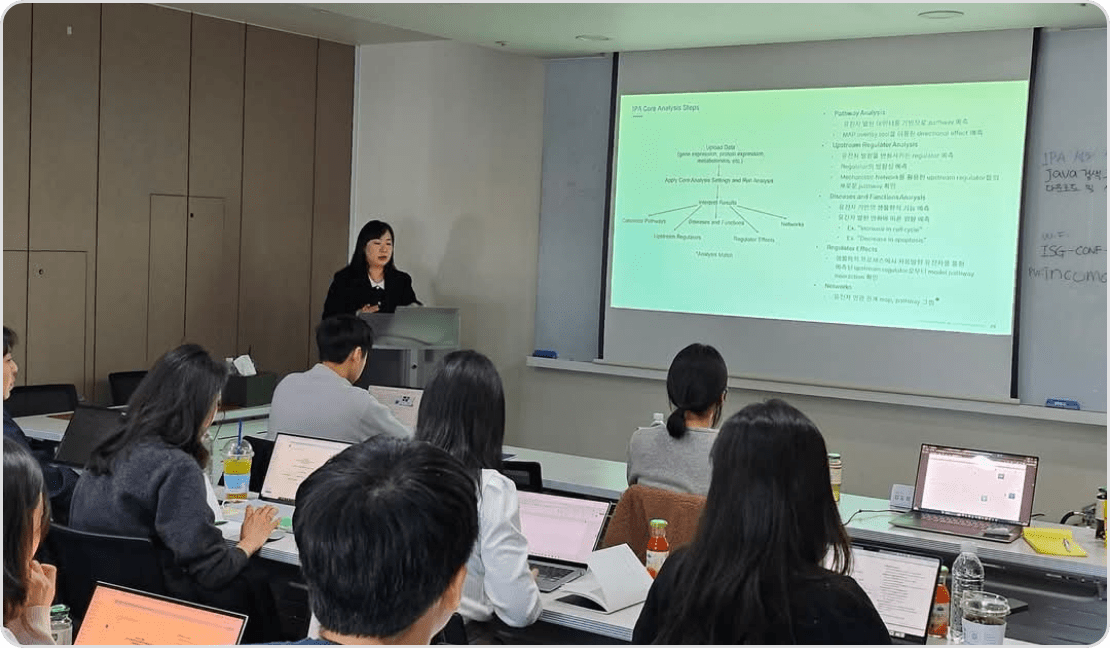
또한, 연구자가 데이터를 직접 분석하고 활용할 수 있도록 생물정보 소프트웨어, 컨설팅, 교육이 결합된 통합 솔루션을 제공합니다.
생물정보, 빅데이터, AI 기술을 융합하여 연구자가 최적의 분석 환경을 구축하고,보다 정확한 바이오 연구를 수행할 수 있도록 지속적으로 지원하겠습니다.
국내·외 최고의 전문가가 빠르고 신뢰도 높은
맞춤형 생물정보 분석 서비스를 제공합니다.
유전체, 전사체, 변이체 분석을 포함한 생명과학 전 분야에 걸쳐 NGS (차세대염기서열분석) 기반의 생물정보 분석 서비스를 제공하며, 고객의 요구와 목적에 맞는 방법론을 제시하고 그 결과로 고객과 대화합니다.
60여 명의 생물정보 전문 인력과 국외 협력 기관이 다년간 구축한 생물정보 분석 파이프라인 및 지식을 통해 고객의 목적에 최적화된 서비스를 제공합니다. 해석하기 어려웠던 기존 데이터부터 특화된 새로운 연구 분야까지 일원화된 서비스를 경험하십시오.
DOMAIN

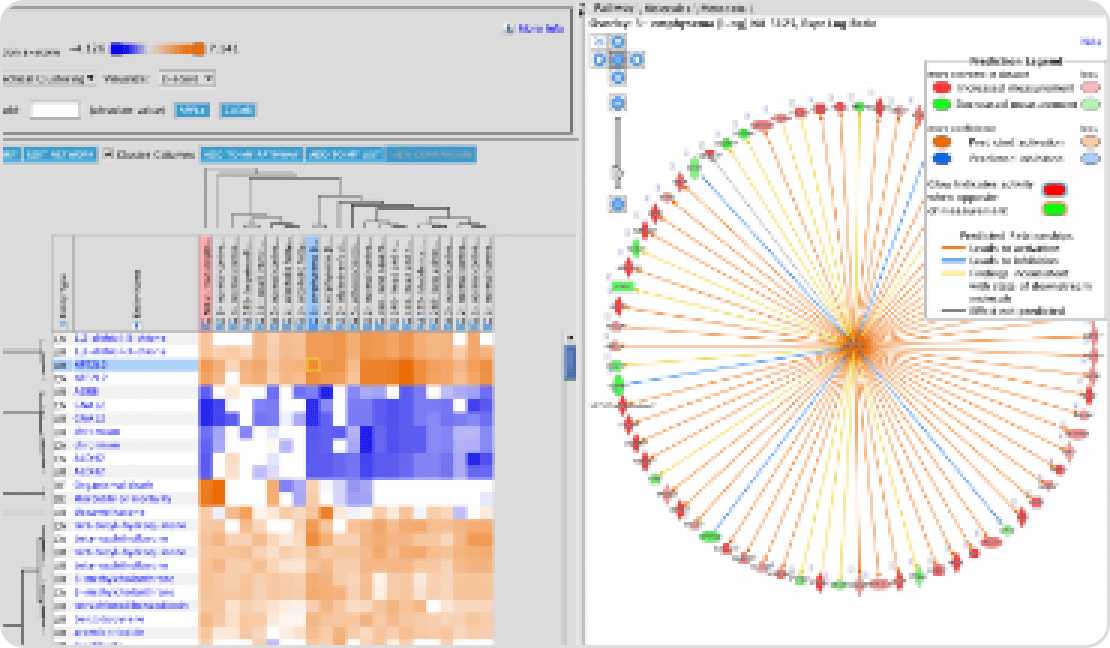
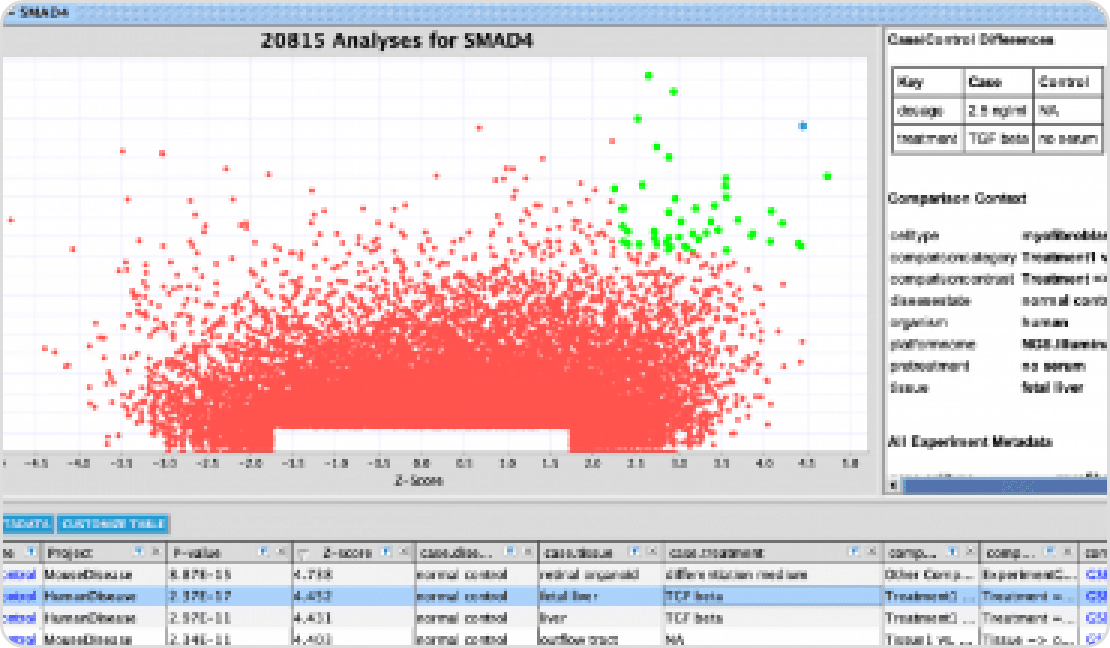
인간 질병 연관 분석, 유전자 네트워크 및 분자기전 연구

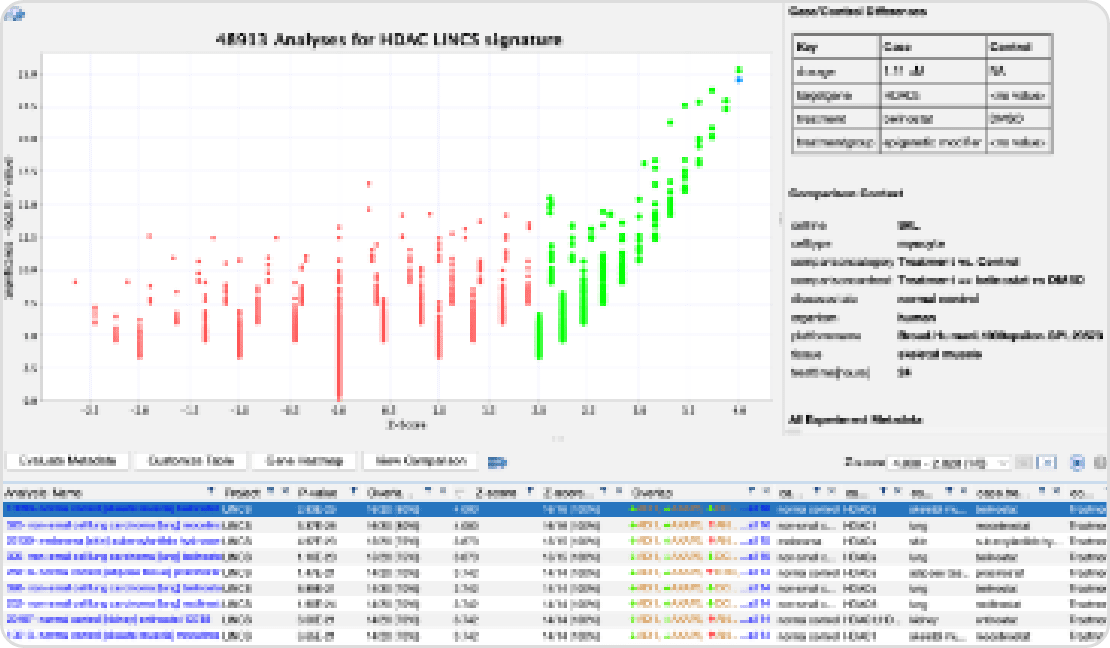
유용 형질 관련 유전자 변이 및 발현 분석

유전체 프로젝트, 유전자 구조 및 기능 분석, 메타게놈 분석
CUSTOMIZED
대표 사례 보기유전자 발현 변화를 분석하여 생물체의 기능적 역할을 규명하는 전사체 데이터 해석 서비스
유전체 변이 탐색을 통해 특정 형질 및 질병 연관성을 분석하는 맞춤형 변이 분석 서비스
고해상도 유전체 서열 분석을 기반으로 유전자 구조 및 기능을 예측하는 유전체 분석 서비스
다양한 생물 종 간 유전체 비교를 통해 공통 및 특이 유전자를 탐색하는 분석 서비스
DNA 메틸화 및 히스톤 변형 데이터를 분석하여 유전자 발현 조절 기작을 연구하는 후성유전체 분석 서비스
개별 세포 수준의 유전체, 전사체, 단백체, 대사체 분석을 통해 세포 간 이질성 이해 및 질병 메커니즘 규명